moabb.paradigms.base.BaseProcessing#
- class moabb.paradigms.base.BaseProcessing(filters: List[Tuple[float, float]], tmin: float = 0.0, tmax: float | None = None, baseline: Tuple[float, float] | None = None, channels: List[str] | None = None, resample: float | None = None, overlap: float | None = None)[source]#
Base Processing.
Please use one of the child classes
- Parameters:
filters (list of list (defaults [[7, 35]])) – bank of bandpass filter to apply.
tmin (float (default 0.0)) – Start time (in second) of the epoch, relative to the dataset specific task interval e.g. tmin = 1 would mean the epoch will start 1 second after the beginning of the task as defined by the dataset.
tmax (float | None, (default None)) – End time (in second) of the epoch, relative to the beginning of the dataset specific task interval. tmax = 5 would mean the epoch will end 5 second after the beginning of the task as defined in the dataset. If None, use the dataset value.
baseline (None | tuple of length 2) – The time interval to consider as “baseline” when applying baseline correction. If None, do not apply baseline correction. If a tuple (a, b), the interval is between a and b (in seconds), including the endpoints. Correction is applied by computing the mean of the baseline period and subtracting it from the data (see mne.Epochs)
channels (list of str | None (default None)) – list of channel to select. If None, use all EEG channels available in the dataset.
resample (float | None (default None)) – If not None, resample the eeg data with the sampling rate provided.
overlap (float | None (default None)) – Overlap percentage (0-100) for the sliding window approach used in pseudo-online evaluation. If None, no overlap is applied. When overlap is used, windows may cross event boundaries; such windows are kept and labeled using a majority vote over the events they cover.
- abstract property datasets#
Property that define the list of compatible datasets.
- get_data(dataset, subjects=None, return_epochs=False, return_raws=False, cache_config=None, postprocess_pipeline=None, process_pipelines=None, additional_metadata: Literal['all'] | list[str] | None = None)[source]#
Return the data for a list of subject.
return the data, labels and a dataframe with metadata. the dataframe will contain at least the following columns
subject : the subject indice
session : the session indice
run : the run indice
- Parameters:
dataset – A dataset instance.
subjects (List of int) – List of subject number
return_epochs (boolean) – This flag specifies whether to return only the data array or the complete processed mne.Epochs
return_raws (boolean) – To return raw files and events, to ensure compatibility with braindecode. Mutually exclusive with return_epochs
cache_config (dict | CacheConfig) – Configuration for caching of datasets. See
moabb.datasets.base.CacheConfigfor details.postprocess_pipeline (Pipeline | None) – Optional pipeline to apply to the data after the preprocessing. This pipeline will either receive
mne.io.BaseRaw,mne.Epochsornp.ndarray()as input, depending on the values ofreturn_epochsandreturn_raws. This pipeline must return annp.ndarray. This pipeline must be “fixed” because it will not be trained, i.e. no call tofitwill be made.process_pipelines (Pipeline | None) – Optional pipeline to apply to the data after the preprocessing. You must set the
return_epochsandreturn_raws` parameters accordingly, i.e., if your custom pipeline returns raw objects, you must also set ``return_raws=True, otherwise you will get unexpected results. Only use it if you know what you are doing.additional_metadata (Literal["all"] | list[str] | None) – Additional metadata to be loaded from the dataset. If None, the default metadata will be loaded containing subject, session and run. If “all”, all columns of the events.tsv file will be loaded. A list of column names can be passed to just select these columns in addition to the three default values mentioned before. This parameter works regardless of the return type (epochs, raws, or array).
- Returns:
X (Union[np.ndarray, mne.Epochs]) – the data that will be used as features for the model Note: if return_epochs=True, this is mne.Epochs if return_epochs=False, this is np.ndarray
labels (np.ndarray) – the labels for training / evaluating the model
metadata (pd.DataFrame) – A dataframe containing the metadata.
- abstract is_valid(dataset)[source]#
Verify the dataset is compatible with the paradigm.
This method is called to verify dataset is compatible with the paradigm.
This method should raise an error if the dataset is not compatible with the paradigm. This is for example the case if the dataset is an ERP dataset for motor imagery paradigm, or if the dataset does not contain any of the required events.
- Parameters:
dataset (dataset instance) – The dataset to verify.
- make_labels_pipeline(dataset, return_epochs=False, return_raws=False)[source]#
Returns the pipeline that extracts the labels from the output of the postprocess_pipeline. Refer to the arguments of
get_data()for more information.
- make_process_pipelines(dataset, return_epochs=False, return_raws=False, postprocess_pipeline=None)[source]#
Create pre-processing pipelines for the data.
Return the pre-processing pipelines corresponding to this paradigm (one per frequency band).
- Parameters:
dataset (BaseDataset) – The dataset instance.
return_epochs (bool, default is False) – Specify if needed to return epochs instead of ndarray.
return_raws (bool, default is False) – Specify if needed to return raws instead of ndarray.
postprocess_pipeline (Pipeline | None, default is None) – Optional pipeline to apply to the data after the preprocessing. This pipeline will either receive
mne.io.BaseRaw,mne.Epochsornp.ndarray()as input, depending on the values ofreturn_epochsandreturn_raws. This pipeline must return annp.ndarray. This pipeline must be “fixed” because it will not be trained, i.e. no call tofitwill be made.
- match_all(datasets: List[BaseDataset], shift=-0.5, channel_merge_strategy: str = 'intersect', ignore=['stim'])[source]#
Initialize this paradigm to match all datasets in parameter:
self.resample is set to match the minimum frequency in all datasets, minus shift. If the frequency is 128 for example, then MNE can return 128 or 129 samples depending on the dataset, even if the length of the epochs is 1s Setting shift=-0.5 solves this particular issue.
self.channels is initialized with the channels which are common to all datasets.
- Parameters:
datasets (List[BaseDataset]) – A dataset instance.
shift (List[BaseDataset]) – Shift the sampling frequency by this value E.g.: if sampling=128 and shift=-0.5, then it returns 127.5 Hz
channel_merge_strategy (str (default: 'intersect')) – Accepts two values: - ‘intersect’: keep only channels common to all datasets - ‘union’: keep all channels from all datasets, removing duplicate
ignore (List[string]) – A list of channels to ignore
..versionadded: – 0.6.0:
- prepare_process(dataset)[source]#
Prepare processing of raw files.
This function allows to set parameter of the paradigm class prior to the preprocessing (process_raw). Does nothing by default and could be overloaded if needed.
- Parameters:
dataset (dataset instance) – The dataset corresponding to the raw file. mainly use to access dataset specific information.
Examples using moabb.paradigms.base.BaseProcessing#

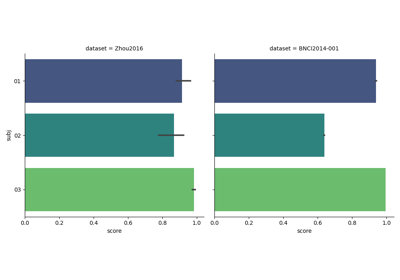
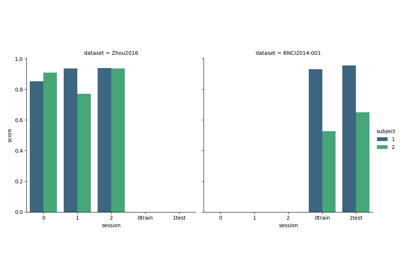
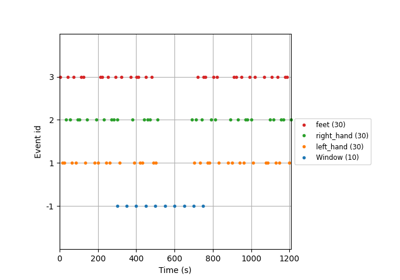
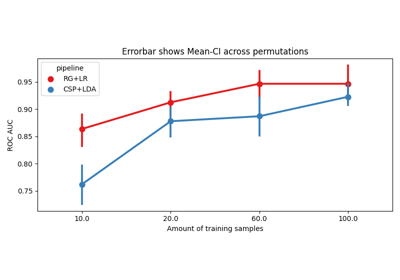
Tutorial 5: Combining Multiple Datasets into a Single Dataset
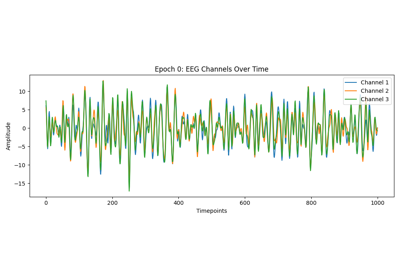
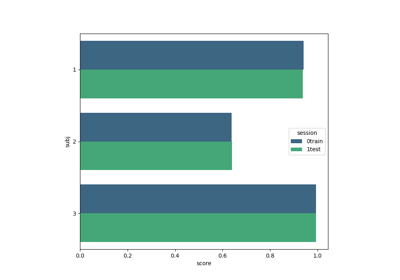
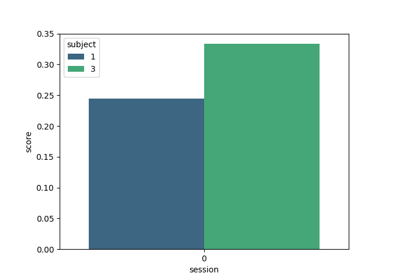
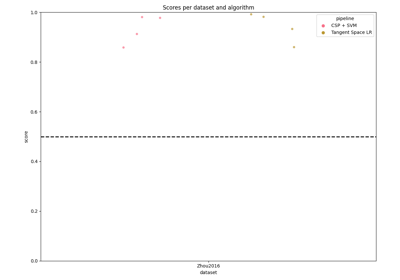
Tutorial: Within-Session Splitting on Real MI Dataset

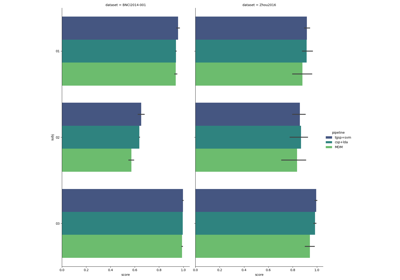
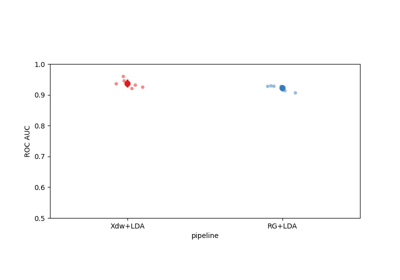
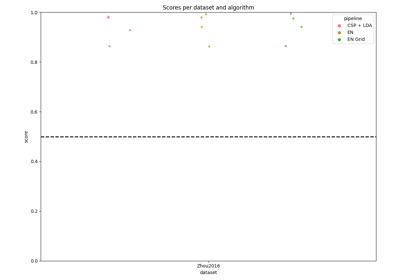
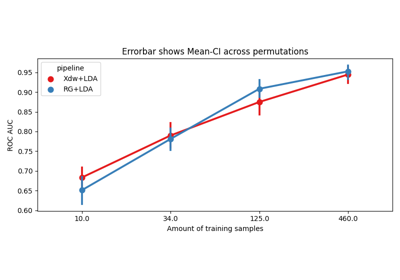
Riemannian Artifact Rejection as a Pre-processing Step

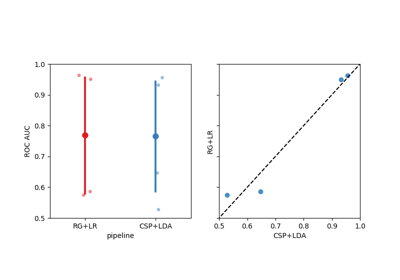
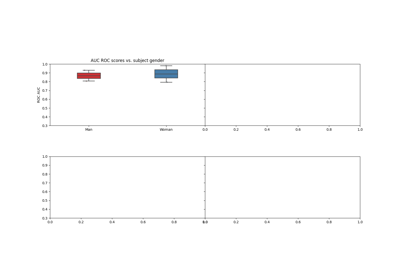
Using X y data (epoched data) instead of continuous signal