moabb.datasets.BNCI2014_009#
- class moabb.datasets.BNCI2014_009(subjects=None, sessions=None)[source]#
Bases:
MNEBNCI[source]Dataset Snapshot
BNCI2014_009
Complete record of P300 evoked potentials recorded with BCI2000 using two different paradigms: P300 Speller (overt attention) and GeoSpell (covert attention). 10 healthy subjects focused on one out of 36 different characters.
P300 / ERP, 2 classes (Target vs NonTarget)
P300 / ERP Code: BNCI2014-009 10 subjects 3 sessions 16 ch 256 Hz 2 classes 16.0 s trials CC BY-NC-ND 4.0Class Labels: Target, NonTarget
Benchmark Context
WithinSessionIncluded in 1 MOABB benchmark table(s). Scores are across available pipelines (WithinSession accuracy).
- ERP/P300 all classes 5 pipelinesMax 93.43 · Median 84.52 · Mean 83.04 · Std 11.79
Citation & Impact
- Paper DOI10.3389/fnhum.2013.00732
- CitationsLoading…
- Public APICrossref | OpenAlex
- Data DOI10.1088/1741-2560/11/3/035008
- MOABB tables1 (WithinSession)
- Page Views30d: 54 · all-time: 553#17 of 151 · Top 12% most viewedUpdated: 2026-03-21 UTC
HED Event TagsHED tagsSource: MOABB BIDS HED annotation mapping.
TargetSensory-eventExperimental-stimulusVisual-presentationTargetNonTargetSensory-eventExperimental-stimulusVisual-presentationNon-targetHED tree view
Tree · Target
├─ Sensory-event ├─ Experimental-stimulus ├─ Visual-presentation └─ Target
Tree · NonTarget
├─ Sensory-event ├─ Experimental-stimulus ├─ Visual-presentation └─ Non-target
Channel SummaryTotal channels16EEG16 (Ag/AgCl)Montage10-10Sampling256 HzReferencelinked earlobesFilterbandpass 0.1-20 HzNotch / line50 HzThis diagram is automatically generated from MOABB metadata. Please consult the original publication to confirm the experimental protocol details.
BNCI 2014-009 P300 dataset.
Dataset from [1].
Dataset Description
This dataset contains EEG data from 10 subjects using a P300 speller system with both grid speller and geo-speller paradigms. This loader includes only the grid speller data.
Participants
10 healthy subjects
Recording Details
Channels: 16 EEG channels
Sampling rate: 256 Hz
Reference: Linked mastoids
References
[1]Riccio, A., Simione, L., Schettini, F., Pizzimenti, A., Inghilleri, M., Belardinelli, M. O., & Mattia, D. (2013). Attention and P300-based BCI performance in people with amyotrophic lateral sclerosis. Frontiers in human neuroscience, 7, 732. https://doi.org/10.3389/fnhum.2013.00732
from moabb.datasets import BNCI2014_009 dataset = BNCI2014_009() data = dataset.get_data(subjects=[1]) print(data[1])
Dataset summary
#Subj
10
#Chan
16
#Trials / class
1440 NT / 288 T
Trials length
0.8 s
Freq
256 Hz
#Sessions
3
Participants
Population: healthy
Age: 26.8 years
BCI experience: experienced
Equipment
Amplifier: g.USBamp
Electrodes: Ag/AgCl
Montage: 10-10
Reference: linked earlobes
Preprocessing
Data state: preprocessed
Bandpass filter: 0.1-20 Hz
Steps: bandpass filtering
Re-reference: linked earlobes
Notes: EEG acquired using g.USBamp amplifier (g.Tec, Austria), digitized at 256 Hz
Data Access
DOI: 10.1088/1741-2560/11/3/035008
Repository: BNCI Horizon
Experimental Protocol
Paradigm: p300
Task type: spelling
Feedback: none
Stimulus: visual_intensification
Notes
Note
BNCI2014_009was previously namedBNCI2014009.BNCI2014009will be removed in version 1.1.Added in version 0.4.0.
- property all_subjects#
Full list of subjects available in this dataset (unfiltered).
- convert_to_bids(path=None, subjects=None, overwrite=False, format='EDF', verbose=None, generate_figures=False)[source]#
Convert the dataset to BIDS format.
Saves the raw EEG data in a BIDS-compliant directory structure. Unlike the caching mechanism (see
CacheConfig), the files produced here do not contain a processing-pipeline hash (desc-<hash>) in their names, making the output a clean, shareable BIDS dataset.- Parameters:
path (str |
Path| None) – Directory under which the BIDS dataset will be written. IfNonethe default MNE data directory is used (same default as the rest of MOABB).subjects (list of int | None) – Subject numbers to convert. If
None, all subjects insubject_listare converted.overwrite (bool) – If
True, existing BIDS files for a subject are removed before saving. Default isFalse.format (str) – The file format for the raw EEG data. Supported values are
"EDF"(default),"BrainVision", and"EEGLAB".verbose (str | None) – Verbosity level forwarded to MNE/MNE-BIDS.
generate_figures (bool) – If
True, generate interactive neural signature HTML figures in{bids_root}/derivatives/neural_signatures/. Requiresplotly(pip install moabb[interactive]). Default isFalse.
- Returns:
bids_root – Path to the root of the written BIDS dataset.
- Return type:
Examples
>>> from moabb.datasets import AlexMI >>> dataset = AlexMI() >>> bids_root = dataset.convert_to_bids(path='/tmp/bids', subjects=[1])
Notes
Use
CacheConfigto configure caching forget_data(). Usemoabb.datasets.bids_interface.get_bids_rootto get the BIDS root path.Added in version 1.5.
- data_path(subject, path=None, force_update=False, update_path=None, verbose=None)[source]#
Get path to local copy of a subject data.
- Parameters:
subject (int) – Number of subject to use
path (None | str) – Location of where to look for the data storing location. If None, the environment variable or config parameter
MNE_DATASETS_(dataset)_PATHis used. If it doesn’t exist, the “~/mne_data” directory is used. If the dataset is not found under the given path, the data will be automatically downloaded to the specified folder.force_update (bool) – Force update of the dataset even if a local copy exists.
update_path (bool | None Deprecated) – If True, set the MNE_DATASETS_(dataset)_PATH in mne-python config to the given path. If None, the user is prompted.
verbose (bool, str, int, or None) – If not None, override default verbose level (see
mne.verbose()).
- Returns:
path – Local path to the given data file. This path is contained inside a list of length one, for compatibility.
- Return type:
- download(subject_list=None, path=None, force_update=False, update_path=None, accept=False, verbose=None)[source]#
Download all data from the dataset.
This function is only useful to download all the dataset at once.
- Parameters:
subject_list (list of int | None) – List of subjects id to download, if None all subjects are downloaded.
path (None | str) – Location of where to look for the data storing location. If None, the environment variable or config parameter
MNE_DATASETS_(dataset)_PATHis used. If it doesn’t exist, the “~/mne_data” directory is used. If the dataset is not found under the given path, the data will be automatically downloaded to the specified folder.force_update (bool) – Force update of the dataset even if a local copy exists.
update_path (bool | None) – If True, set the MNE_DATASETS_(dataset)_PATH in mne-python config to the given path. If None, the user is prompted.
accept (bool) – Accept licence term to download the data, if any. Default: False
verbose (bool, str, int, or None) – If not None, override default verbose level (see
mne.verbose()).
- get_additional_metadata(subject: str, session: str, run: str)[source]#
Load additional metadata for a specific subject, session, and run.
This method is intended to be overridden by subclasses to provide additional metadata specific to the dataset. The metadata is typically loaded from an events.tsv file or similar data source.
- Parameters:
- Returns:
A DataFrame containing the additional metadata if available, otherwise None.
- Return type:
None |
pandas.DataFrame
- get_block_repetition(paradigm, subjects, block_list, repetition_list)[source]#
Select data for all provided subjects, blocks and repetitions.
subject -> session -> run -> block -> repetition
See also
- get_data(subjects=None, cache_config=None, process_pipeline=None)[source]#
Return the data corresponding to a list of subjects.
The returned data is a dictionary with the following structure:
data = {'subject_id' : {'session_id': {'run_id': run} } }
subjects are on top, then we have sessions, then runs. A sessions is a recording done in a single day, without removing the EEG cap. A session is constitued of at least one run. A run is a single contiguous recording. Some dataset break session in multiple runs.
Processing steps can optionally be applied to the data using the
*_pipelinearguments. These pipelines are applied in the following order:raw_pipeline->epochs_pipeline->array_pipeline. If a*_pipelineargument isNone, the step will be skipped. Therefore, thearray_pipelinemay either receive amne.io.Rawor amne.Epochsobject as input depending on whetherepochs_pipelineisNoneor not.- Parameters:
subjects (List of int) – List of subject number
cache_config (dict |
CacheConfig) – Configuration for caching of datasets. SeeCacheConfigfor details.process_pipeline (
sklearn.pipeline.Pipeline| None) – Optional processing pipeline to apply to the data. To generate an adequate pipeline, we recommend usingmoabb.make_process_pipelines(). This pipeline will receivemne.io.BaseRawobjects. The steps names of this pipeline should be elements ofStepType. According to their name, the steps should either return amne.io.BaseRaw, amne.Epochs, or anumpy.ndarray. This pipeline must be “fixed” because it will not be trained, i.e. no call tofitwill be made.
- Returns:
data – dict containing the raw data
- Return type:
Dict
- property metadata[source]#
Return structured metadata for this dataset.
Returns the DatasetMetadata object from the centralized catalog, or None if metadata is not available for this dataset.
- Returns:
The metadata object containing acquisition parameters, participant demographics, experiment details, and documentation. Returns None if no metadata is registered for this dataset.
- Return type:
DatasetMetadata| None
Examples
>>> from moabb.datasets import BNCI2014_001 >>> dataset = BNCI2014_001() >>> dataset.metadata.participants.n_subjects 9 >>> dataset.metadata.acquisition.sampling_rate 250.0
Examples using moabb.datasets.BNCI2014_009#

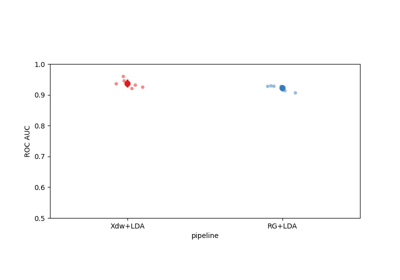
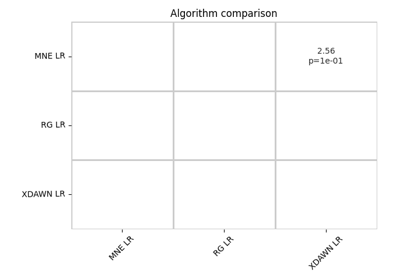
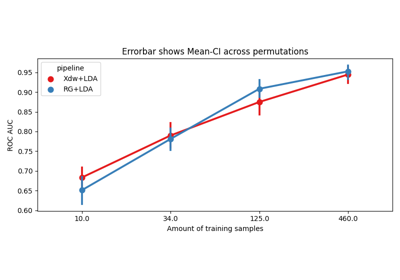
Riemannian Artifact Rejection as a Pre-processing Step