moabb.datasets.Dreyer2023A#
- class moabb.datasets.Dreyer2023A(subjects=None, sessions=None, *, return_all_modalities=False)[source]#
Bases:
_Dreyer2023Base[source]Dataset Snapshot
Dreyer2023A
A large EEG database with users' profile information for motor imagery brain-computer interface research. Contains electroencephalographic signals from 87 human participants, collected during a single day of brain-computer interface (BCI) experiments, organized into 3 datasets (A, B, and C) that were all recorded using the same protocol: right and left hand motor imagery (MI).
Motor Imagery, 2 classes (left_hand vs right_hand)
Class Labels: left_hand, right_hand
Citation & Impact
- Paper DOI10.1038/s41597-023-02445-z
- CitationsLoading…
- Public APICrossref | OpenAlex
- Page Views30d: 33 · all-time: 215#39 of 152 · Top 26% most viewedUpdated: 2026-03-30 UTC
HED Event TagsHED tagsSource: MOABB BIDS HED annotation mapping.
left_handSensory-eventAgent-actionright_handSensory-eventAgent-actionHED tree view
Tree · left_hand
├─ Sensory-event │ ├─ Experimental-stimulus │ └─ Visual-presentation └─ Agent-action └─ Imagine ├─ Move └─ Left └─ HandTree · right_hand
├─ Sensory-event │ ├─ Experimental-stimulus │ └─ Visual-presentation └─ Agent-action └─ Imagine ├─ Move └─ Right └─ HandChannel SummaryTotal channels27EEG27 (active electrodes)EOG3EMG2Montage10-20Sampling512 HzReferenceleft earlobeFilternone (raw signals recorded without hardware filters)Notch / line50 HzThis diagram is automatically generated from MOABB metadata. Please consult the original publication to confirm the experimental protocol details.
Class for Dreyer2023A dataset management. MI dataset.
Dataset description
“A large EEG database with users’ profile information for motor imagery Brain-Computer Interface research” [1] [2]
- Data collectors:
Appriou Aurélien; Caselli Damien; Benaroch Camille; Yamamoto Sayu Maria; Roc Aline; Lotte Fabien; Dreyer Pauline; Pillette Léa
- Data manager:
Dreyer Pauline
- Project leader:
Lotte Fabien
- Project members:
Rimbert Sébastien; Monseigne Thibaut
Dataset Dreyer2023A contains EEG, EOG and EMG signals recorded on 60 healthy subjects performing Left-Right Motor Imagery experiments (29 women, age 19-59, M = 29, SD = 9.32) [1]. Experiments were conducted by six experimenters. In addition, for each recording the following pieces of information are provided: subject’s demographic, personality and cognitive profiles, the OpenViBE experimental instructions and codes, and experimenter’s gender.
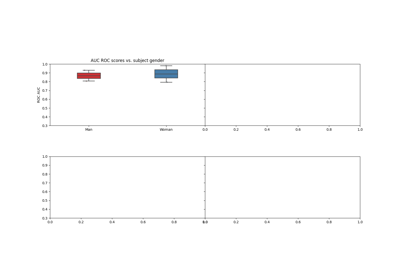
The experiment is designed for the investigation of the impact of the participant’s and experimenter’s gender on MI BCI performance [1].
A recording contains open and closed eyes baseline recordings and 6 runs of the MI experiments. First 2 runs (acquisition runs) were used to train system and the following 4 runs (training runs) to train the participant. Each run contained 40 trials [1].
- Each trial was recorded as follows [1]:
t=0.00s cross displayed on screen
t=2.00s acoustic signal announced appearance of a red arrow
t=3.00s a red arrow appears (subject starts to perform task)
t=4.25s the red arrow disappears
t=4.25s the feedback on performance is given in form of a blue bar with update frequency of 16 Hz
t=8.00s cross turns off (subject stops to perform task)
- EEG signals [1]:
recorded with 27 electrodes, namely: Fz, FCz, Cz, CPz, Pz, C1, C3, C5, C2, C4, C6, F4, FC2, FC4, FC6, CP2, CP4, CP6, P4, F3, FC1, FC3, FC5, CP1, CP3, CP5, P3 (10-20 system), referenced to the left earlobe.
- EOG signals [1]:
recorded with 3 electrodes, namely: EOG1, EOG2, EOG3 placed below, above and on the side of one eye.
- EMG signals [1]:
recorded with 2 electrodes, namely: EMGg, EMGd placed 2.5cm below the skinfold on each wrist.
- Demographic and biosocial information includes:
gender, birth year, laterality
vision, vision assistance
familiarity to cognitive science or neurology, level of education
physical activity, meditation
attentional, neurological, psychiatrics symptoms
- Personality and the cognitive profile [1]:
evaluated via 5th edition of the 16 Personality Factors (16PF5) test
and mental rotation test
index of learning style
- Pre and post experiment questionnaires [1]:
evaluation of pre and post mood, mindfulness and motivational states
- The online OpenViBE BCI classification performance [1]:
only performance measure used to give the feedback to the participants
Subject 59 contains only 4 runs
References
[1] (1,2,3,4,5,6,7,8,9,10,11)Pillette, L., Roc, A., N’kaoua, B., & Lotte, F. (2021). Experimenters’ influence on mental-imagery based brain-computer interface user training. International Journal of Human-Computer Studies, 149, 102603.
[2]Benaroch, C., Yamamoto, M. S., Roc, A., Dreyer, P., Jeunet, C., & Lotte, F. (2022). When should MI-BCI feature optimization include prior knowledge, and which one?. Brain-Computer Interfaces, 9(2), 115-128.
from moabb.datasets import Dreyer2023A dataset = Dreyer2023A() data = dataset.get_data(subjects=[1]) print(data[1])
Dataset summary
#Subj
60
#Chan
27
#Classes
2
#Trials / class
20
Trials length
5 s
Freq
512 Hz
#Sessions
1
#Runs
6
Total_trials
14400
- __init__(subjects=None, sessions=None, *, return_all_modalities=False)[source]#
Initialize function for the BaseDataset.
- property all_subjects#
Full list of subjects available in this dataset (unfiltered).
- convert_to_bids(path=None, subjects=None, overwrite=False, format='EDF', verbose=None, generate_figures=False)[source]#
Convert the dataset to BIDS format.
Saves the raw EEG data in a BIDS-compliant directory structure. Unlike the caching mechanism (see
CacheConfig), the files produced here do not contain a processing-pipeline hash (desc-<hash>) in their names, making the output a clean, shareable BIDS dataset.- Parameters:
path (str |
Path| None) – Directory under which the BIDS dataset will be written. IfNonethe default MNE data directory is used (same default as the rest of MOABB).subjects (list of int | None) – Subject numbers to convert. If
None, all subjects insubject_listare converted.overwrite (bool) – If
True, existing BIDS files for a subject are removed before saving. Default isFalse.format (str) – The file format for the raw EEG data. Supported values are
"EDF"(default),"BrainVision", and"EEGLAB".verbose (str | None) – Verbosity level forwarded to MNE/MNE-BIDS.
generate_figures (bool) – If
True, generate interactive neural signature HTML figures in{bids_root}/derivatives/neural_signatures/. Requiresplotly(pip install moabb[interactive]). Default isFalse.
- Returns:
bids_root – Path to the root of the written BIDS dataset.
- Return type:
Examples
>>> from moabb.datasets import AlexMI >>> dataset = AlexMI() >>> bids_root = dataset.convert_to_bids(path='/tmp/bids', subjects=[1])
Notes
Use
CacheConfigto configure caching forget_data(). Usemoabb.datasets.bids_interface.get_bids_rootto get the BIDS root path.Added in version 1.5.
- data_path(subject, path=None, force_update=False, update_path=None, verbose=None)[source]#
Return the data BIDS paths of a single subject.
- Parameters:
subject (int) – The subject number to fetch data for.
path (None | str) – Location of where to look for the data storing location. If None, the environment variable or config parameter MNE_(dataset) is used. If it doesn’t exist, the “~/mne_data” directory is used. If the dataset is not found under the given path, the data will be automatically downloaded to the specified folder.
force_update (bool) – Force update of the dataset even if a local copy exists.
update_path (bool | None) – If True, set the MNE_DATASETS_(dataset)_PATH in mne-python config to the given path. If None, the user is prompted.
verbose (bool, str, int, or None) – If not None, override default verbose level (see mne.verbose()).
- Returns:
A list containing the BIDSPath object for the subject’s data file.
- Return type:
- download(subject_list=None, path=None, force_update=False, update_path=None, accept=False, verbose=None)[source]#
Download all data from the dataset.
This function is only useful to download all the dataset at once.
- Parameters:
subject_list (list of int | None) – List of subjects id to download, if None all subjects are downloaded.
path (None | str) – Location of where to look for the data storing location. If None, the environment variable or config parameter
MNE_DATASETS_(dataset)_PATHis used. If it doesn’t exist, the “~/mne_data” directory is used. If the dataset is not found under the given path, the data will be automatically downloaded to the specified folder.force_update (bool) – Force update of the dataset even if a local copy exists.
update_path (bool | None) – If True, set the MNE_DATASETS_(dataset)_PATH in mne-python config to the given path. If None, the user is prompted.
accept (bool) – Accept licence term to download the data, if any. Default: False
verbose (bool, str, int, or None) – If not None, override default verbose level (see
mne.verbose()).
- get_additional_metadata(subject: str, session: str, run: str)[source]#
Load additional metadata for a specific subject, session, and run.
This method is intended to be overridden by subclasses to provide additional metadata specific to the dataset. The metadata is typically loaded from an events.tsv file or similar data source.
- Parameters:
- Returns:
A DataFrame containing the additional metadata if available, otherwise None.
- Return type:
None |
pandas.DataFrame
- get_block_repetition(paradigm, subjects, block_list, repetition_list)[source]#
Select data for all provided subjects, blocks and repetitions.
subject -> session -> run -> block -> repetition
See also
- get_data(subjects=None, cache_config=None, process_pipeline=None)[source]#
Return the data corresponding to a list of subjects.
The returned data is a dictionary with the following structure:
data = {'subject_id' : {'session_id': {'run_id': run} } }
subjects are on top, then we have sessions, then runs. A sessions is a recording done in a single day, without removing the EEG cap. A session is constitued of at least one run. A run is a single contiguous recording. Some dataset break session in multiple runs.
Processing steps can optionally be applied to the data using the
*_pipelinearguments. These pipelines are applied in the following order:raw_pipeline->epochs_pipeline->array_pipeline. If a*_pipelineargument isNone, the step will be skipped. Therefore, thearray_pipelinemay either receive amne.io.Rawor amne.Epochsobject as input depending on whetherepochs_pipelineisNoneor not.- Parameters:
subjects (List of int) – List of subject number
cache_config (dict |
CacheConfig) – Configuration for caching of datasets. SeeCacheConfigfor details.process_pipeline (
sklearn.pipeline.Pipeline| None) – Optional processing pipeline to apply to the data. To generate an adequate pipeline, we recommend usingmoabb.make_process_pipelines(). This pipeline will receivemne.io.BaseRawobjects. The steps names of this pipeline should be elements ofStepType. According to their name, the steps should either return amne.io.BaseRaw, amne.Epochs, or anumpy.ndarray. This pipeline must be “fixed” because it will not be trained, i.e. no call tofitwill be made.
- Returns:
data – dict containing the raw data
- Return type:
Dict
- get_subject_info(path=None)[source]#
Return the demographic information of the subjects.
- Returns:
A DataFrame containing the demographic information of the subjects.
- Return type:
- property metadata[source]#
Return structured metadata for this dataset.
Returns the DatasetMetadata object from the centralized catalog, or None if metadata is not available for this dataset.
- Returns:
The metadata object containing acquisition parameters, participant demographics, experiment details, and documentation. Returns None if no metadata is registered for this dataset.
- Return type:
DatasetMetadata| None
Examples
>>> from moabb.datasets import BNCI2014_001 >>> dataset = BNCI2014_001() >>> dataset.metadata.participants.n_subjects 9 >>> dataset.metadata.acquisition.sampling_rate 250.0