Note
Go to the end to download the full example code.
Within Session P300#
This example shows how to perform a within session analysis on three different P300 datasets.
We will compare two pipelines :
Riemannian geometry
XDAWN with Linear Discriminant Analysis
We will use the P300 paradigm, which uses the AUC as metric.
# Authors: Pedro Rodrigues <pedro.rodrigues01@gmail.com>
#
# License: BSD (3-clause)
import warnings
import matplotlib.pyplot as plt
from mne.decoding import Vectorizer
from pyriemann.estimation import Xdawn, XdawnCovariances
from pyriemann.tangentspace import TangentSpace
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis as LDA
from sklearn.pipeline import make_pipeline
import moabb
import moabb.analysis.plotting as moabb_plt
from moabb.analysis.chance_level import chance_by_chance
from moabb.datasets import BNCI2014_009
from moabb.evaluations import WithinSessionEvaluation
from moabb.paradigms import P300
getting rid of the warnings about the future
warnings.simplefilter(action="ignore", category=FutureWarning)
warnings.simplefilter(action="ignore", category=RuntimeWarning)
moabb.set_log_level("info")
Create Pipelines#
Pipelines must be a dict of sklearn pipeline transformer.
pipelines = {}
We have to do this because the classes are called ‘Target’ and ‘NonTarget’ but the evaluation function uses a LabelEncoder, transforming them to 0 and 1
labels_dict = {"Target": 1, "NonTarget": 0}
pipelines["RG+LDA"] = make_pipeline(
XdawnCovariances(
nfilter=2, classes=[labels_dict["Target"]], estimator="lwf", xdawn_estimator="scm"
),
TangentSpace(),
LDA(solver="lsqr", shrinkage="auto"),
)
pipelines["Xdw+LDA"] = make_pipeline(
Xdawn(nfilter=2, estimator="scm"), Vectorizer(), LDA(solver="lsqr", shrinkage="auto")
)
Evaluation#
We define the paradigm (P300) and use all three datasets available for it. The evaluation will return a DataFrame containing a single AUC score for each subject / session of the dataset, and for each pipeline.
Results are saved into the database, so that if you add a new pipeline, it will not run again the evaluation unless a parameter has changed. Results can be overwritten if necessary.
paradigm = P300(resample=128)
dataset = BNCI2014_009()
dataset.subject_list = dataset.subject_list[:2]
datasets = [dataset]
overwrite = True # set to True if we want to overwrite cached results
evaluation = WithinSessionEvaluation(
paradigm=paradigm, datasets=datasets, suffix="examples", overwrite=overwrite
)
results = evaluation.process(pipelines)
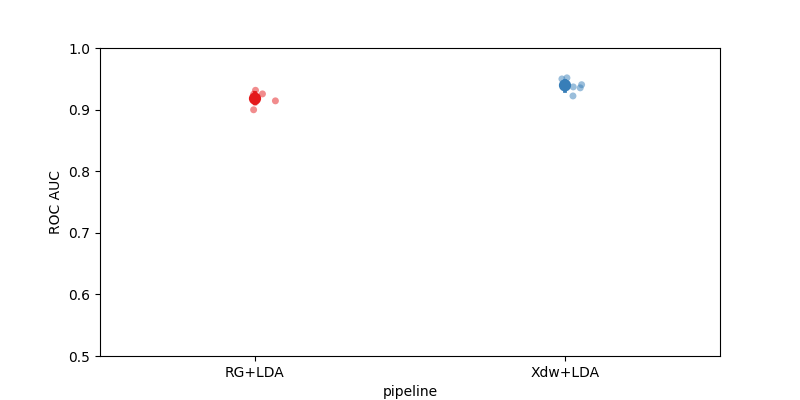
Plot Results#
Here we plot the results using the MOABB score plot with chance level annotations. The P300 paradigm has 2 classes (Target / NonTarget) with a theoretical chance level of 50%.
chance_levels = chance_by_chance(results, alpha=[0.05, 0.01])
fig, _ = moabb_plt.score_plot(results, chance_level=chance_levels)
plt.show()
Total running time of the script: (3 minutes 18.246 seconds)