Note
Go to the end to download the full example code.
Select Electrodes and Resampling#
Within paradigm, it is possible to restrict analysis only to a subset of electrodes and to resample to a specific sampling rate. There is also a utility function to select common electrodes shared between datasets. This tutorial demonstrates how to use this functionality.
# Authors: Sylvain Chevallier <sylvain.chevallier@uvsq.fr>
#
# License: BSD (3-clause)
import matplotlib.pyplot as plt
from mne.decoding import CSP
from pyriemann.estimation import Covariances
from pyriemann.tangentspace import TangentSpace
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis as LDA
from sklearn.linear_model import LogisticRegression as LR
from sklearn.pipeline import make_pipeline
import moabb.analysis.plotting as moabb_plt
from moabb.analysis.chance_level import chance_by_chance
from moabb.datasets import BNCI2014_001, Zhou2016
from moabb.datasets.utils import find_intersecting_channels
from moabb.evaluations import WithinSessionEvaluation
from moabb.paradigms import LeftRightImagery
Datasets#
Load 2 subjects of BNCI 2014-004 and Zhou2016 datasets, with 2 sessions each
subj = [1, 2]
datasets = [Zhou2016(), BNCI2014_001()]
for d in datasets:
d.subject_list = subj
Paradigm#
Restrict further analysis to specified channels, here C3, C4, and Cz. Also, use a specific resampling. In this example, all datasets are set to 200 Hz.
paradigm = LeftRightImagery(channels=["C3", "C4", "Cz"], resample=200.0)
Evaluation#
The evaluation is conducted on with CSP+LDA, only on the 3 electrodes, with a sampling rate of 200 Hz.
evaluation = WithinSessionEvaluation(paradigm=paradigm, datasets=datasets)
csp_lda = make_pipeline(CSP(n_components=2), LDA())
ts_lr = make_pipeline(
Covariances(estimator="oas"), TangentSpace(metric="riemann"), LR(C=1.0)
)
results = evaluation.process({"csp+lda": csp_lda, "ts+lr": ts_lr})
print(results.head())
score time samples ... dataset pipeline codecarbon_task_name
0 0.938000 0.019291 100.0 ... Zhou2016 csp+lda
1 0.853788 0.018971 119.0 ... Zhou2016 csp+lda
2 0.892000 0.017121 100.0 ... Zhou2016 csp+lda
3 0.918000 0.015647 100.0 ... Zhou2016 csp+lda
4 0.920000 0.015703 100.0 ... Zhou2016 csp+lda
[5 rows x 13 columns]
Electrode Selection#
It is possible to select the electrodes that are shared by all datasets using the find_intersecting_channels function. Datasets that have 0 overlap with others are discarded. It returns the set of common channels, as well as the list of datasets with valid channels.
electrodes, datasets = find_intersecting_channels(datasets)
evaluation = WithinSessionEvaluation(
paradigm=paradigm, datasets=datasets, overwrite=True, suffix="resample"
)
results = evaluation.process({"csp+lda": csp_lda, "ts+lr": ts_lr})
print(results.head())
score time samples ... dataset pipeline codecarbon_task_name
0 0.962000 0.014121 100.0 ... Zhou2016 csp+lda
1 0.896000 0.014064 100.0 ... Zhou2016 csp+lda
2 0.860732 0.015478 119.0 ... Zhou2016 csp+lda
3 0.934000 0.013948 100.0 ... Zhou2016 csp+lda
4 0.834568 0.013213 90.0 ... Zhou2016 csp+lda
[5 rows x 13 columns]
Plot Results#
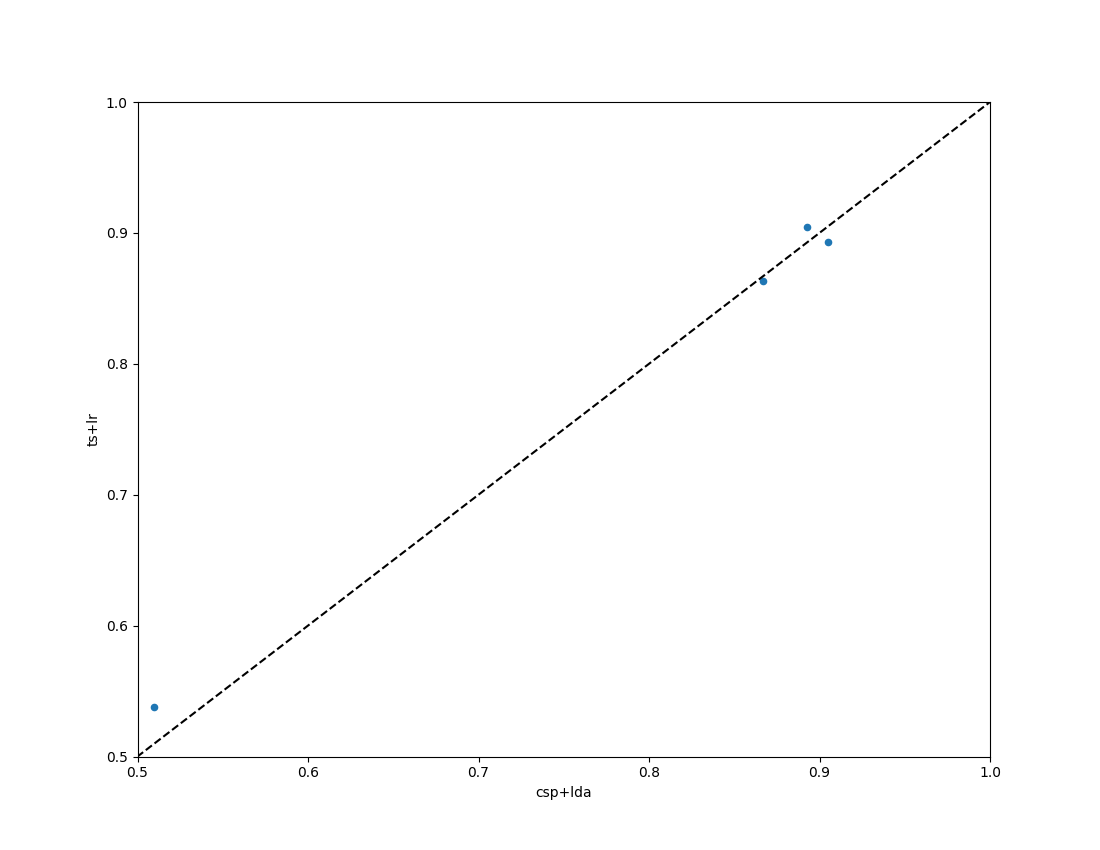
Compare the obtained results with the two pipelines, CSP+LDA and logistic regression computed in the tangent space of the covariance matrices.
chance_levels = chance_by_chance(results, alpha=[0.05, 0.01])
fig = moabb_plt.paired_plot(results, "csp+lda", "ts+lr", chance_level=chance_levels)
plt.show()
Total running time of the script: (10 minutes 38.140 seconds)