moabb.evaluations.base.BaseEvaluation#
- class moabb.evaluations.base.BaseEvaluation(paradigm: BaseParadigm, datasets: list[BaseDataset] | None = None, random_state: int | None = None, n_jobs: int = 1, overwrite: bool = False, error_score: str | float = 'raise', suffix: str = '', hdf5_path: str | None = None, additional_columns: list[str] | None = None, return_epochs: bool = False, return_raws: bool = False, mne_labels: bool = False, n_splits: int | None = None, cv_class: type | None = None, cv_kwargs: dict | None = None, save_model: bool = False, cache_config: CacheConfig | None = None, optuna: bool = False, time_out: int = 900, verbose: bool | str | int | None = None, codecarbon_config: dict | None = None)[source]#
Base class that defines necessary operations for an evaluation. Evaluations determine what the train and test sets are and can implement additional data preprocessing steps for more complicated algorithms.
- Parameters:
paradigm (
BaseParadigm) – The paradigm to use.datasets (list of
BaseDataset) – The list of dataset to run the evaluation. If none, the list of compatible dataset will be retrieved from the paradigm instance.random_state (int or None) – If not None, can guarantee same seed for shuffling examples. Defaults to
None.n_jobs (int) – Number of jobs for fitting of pipeline. Defaults to
1.overwrite (bool) – If true, overwrite the results. Defaults to
False.error_score (str or float) – Value to assign to the score if an error occurs in estimator fitting. If set to
’raise’, the error is raised. Defaults to"raise".suffix (str) – Suffix for the results file.
hdf5_path (str) – Specific path for storing the results.
additional_columns (None) – Adding information to results.
return_epochs (bool) – Use MNE epoch to train pipelines. Defaults to
False.return_raws (bool) – Use MNE raw to train pipelines. Defaults to
False.mne_labels (bool) – If returning MNE epoch, use original dataset label if True. Defaults to
False.n_splits (int or None) – Number of splits for cross-validation. If None, the number of splits is equal to the number of subjects. Defaults to
None.cv_class (type or None) – Optional cross-validation class to override the evaluation’s default splitter behavior. Defaults to
None.cv_kwargs (dict or None) – Keyword arguments passed to cv_class when constructing the splitter. Defaults to
None.save_model (bool) – Save model after training, for each fold of cross-validation if needed. Defaults to
False.cache_config (
CacheConfigor None) – Configuration for caching of datasets. Seemoabb.datasets.base.CacheConfigfor details. Defaults toNone.optuna (bool) – If optuna is enable it will change the GridSearch to a RandomizedGridSearch with 15 minutes of cut off time. This option is compatible with list of entries of type None, bool, int, float and string. Defaults to
False.time_out (int) – Cut off time for the optuna search expressed in seconds. Only used with optuna equal to True. Defaults to
60*15(15 minutes).verbose (bool, str, int, or None) – If not None, override the default MOABB logging level used by this evaluation (see
moabb.utils.verbosefor more information on how this is handled). If used, it should be passed as a keyword-argument only. Defaults toNone.codecarbon_config (dict or None) – Allow CodeCarbon script level configurations. Can use combination of CodeCarbon environment variable and configuration files. See CodeCarbon developer documentation for more information. Defaults to
dict(save_to_file=False, log_level="error").
Notes
Added in version 1.1.0: n_splits, save_model, cache_config parameters.
Added in version 1.1.1: optuna, time_out parameters.
Added in version 1.5: verbose parameter.
- abstract evaluate(dataset: BaseDataset, pipelines: dict[str, BaseEstimator], param_grid: dict | None, process_pipeline: BaseEstimator, postprocess_pipeline: BaseEstimator | None = None)[source]#
Evaluate results on a single dataset.
This method return a generator. each results item is a dict with the following conversion:
res = {'time': Duration of the training , 'dataset': dataset id, 'subject': subject id, 'session': session id, 'score': score, 'n_samples': number of training examples, 'n_channels': number of channel, 'pipeline': pipeline name}
- abstract is_valid(dataset: BaseDataset) bool[source]#
Verify the dataset is compatible with evaluation.
This method is called to verify dataset given in the constructor are compatible with the evaluation context.
This method should return false if the dataset does not match the evaluation. This is for example the case if the dataset does not contain enough session for a cross-session eval.
- Parameters:
dataset (
BaseDataset) – The dataset to verify.
- process(pipelines: dict[str, BaseEstimator], param_grid: dict[str, dict] | None = None, postprocess_pipeline: BaseEstimator | None = None)[source]#
Runs all pipelines on all datasets.
This function will apply all provided pipelines and return a dataframe containing the results of the evaluation.
- Parameters:
pipelines (dict) – A dict of pipeline instances containing the sklearn pipelines to evaluate.
param_grid (dict of str) – The key of the dictionary must be the same as the associated pipeline.
postprocess_pipeline (
sklearn.pipeline.Pipeline| None) – Optional pipeline to apply to the data after the preprocessing. This pipeline will either receivemne.io.BaseRaw,mne.Epochsornumpy.ndarrayas input, depending on the values ofreturn_epochsandreturn_raws. This pipeline must return anumpy.ndarray. This pipeline must be “fixed” because it will not be trained, i.e. no call tofitwill be made.
- Returns:
results – A dataframe containing the results.
- Return type:
Examples using moabb.evaluations.base.BaseEvaluation#

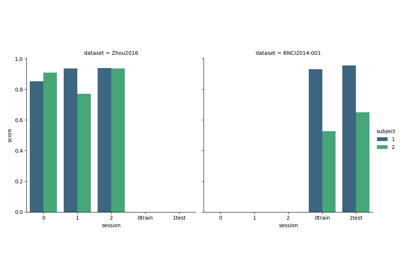
Tutorial 5: Combining Multiple Datasets into a Single Dataset

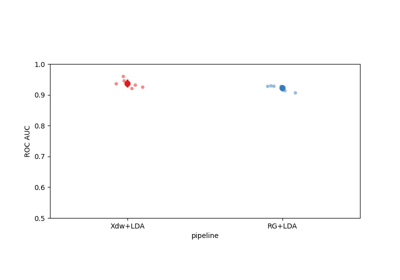
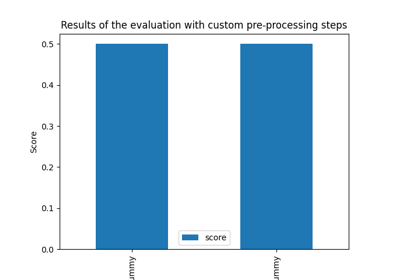
Riemannian Artifact Rejection as a Pre-processing Step

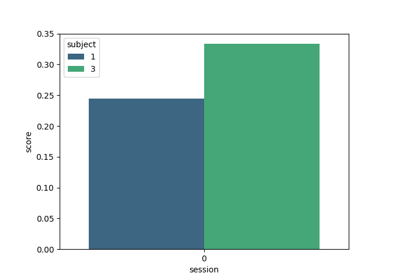
Using X y data (epoched data) instead of continuous signal